Clonogenic (or colony formation) assay is an in vitro cell survival assay that evaluates the ability of a single cell to form a colony. The assay has been widely used to test the efficacy of cancer drugs, gene therapies, and other cytotoxic agents; however, the manual examination and quantification of formed colonies can be a time-consuming and laborious task. An automated approach allows you increase clonogenic assay throughput and reproducibility and gain a kinetic insight into the effect of drugs on cell growth.
Evaluation of chemotherapies using clonogenic assays
-
Automatically measure the number of colonies in a well>
-
Determine a clonogenic assay tipping point by size and number>
-
Measure dose-dependent effects on colony growth >
-
Application Note: Quantifying chemotoxicity on cancer cell colonies>
-
AI-driven quantification of colony formation>
Purpose: To demonstrate whole-well quantification of colony number following anti-cancer (Dox) treatment. Plated at low densities, a whole-well quantification is necessary to ensure accurate assessment of the surviving fraction.
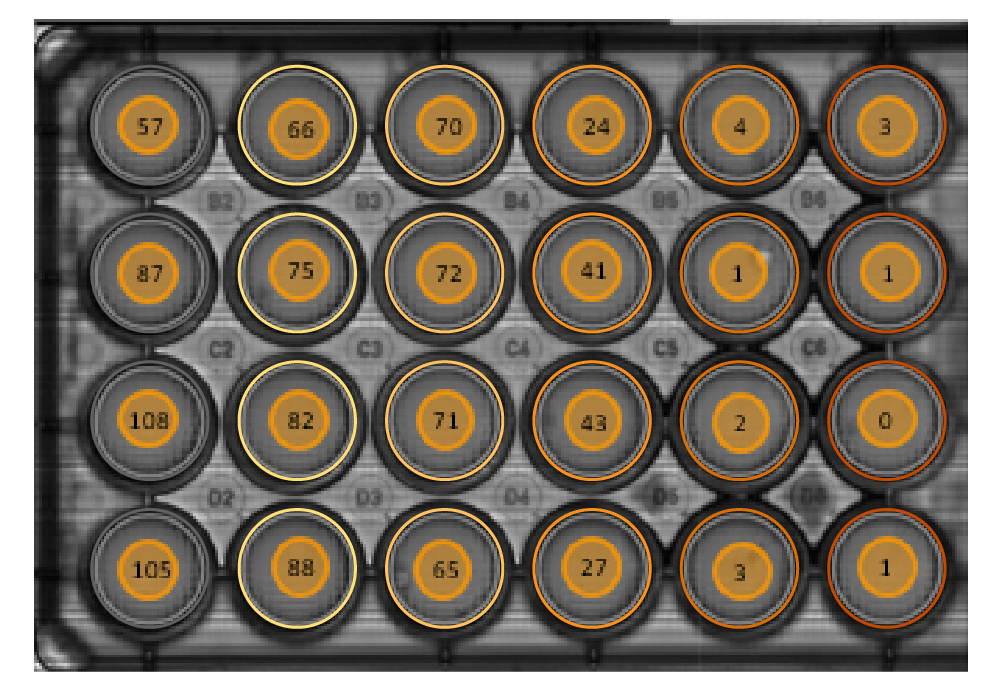
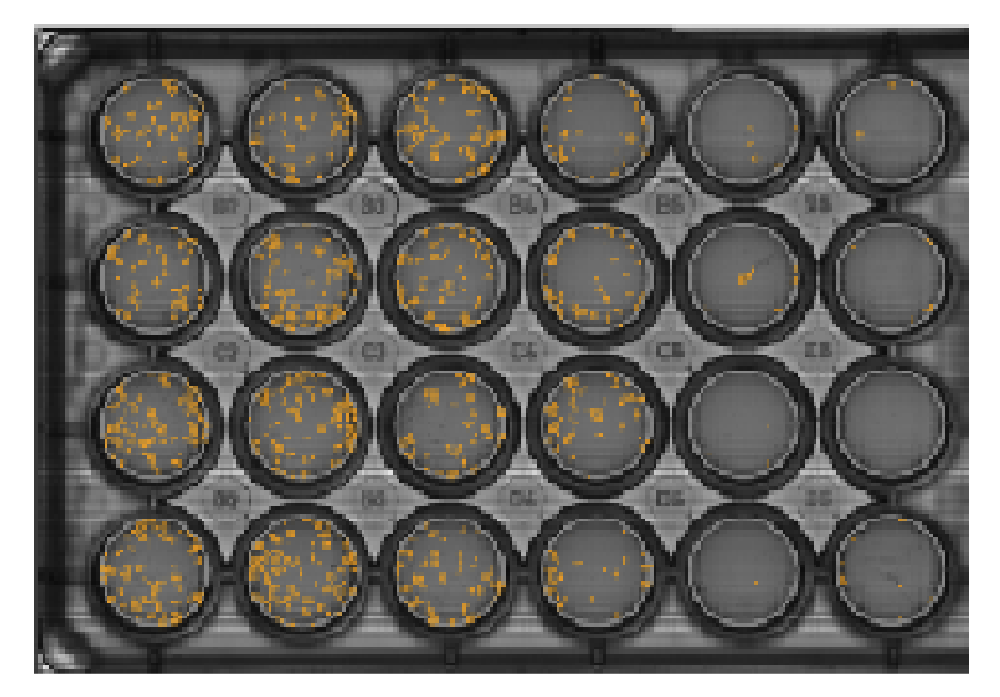


The Clonogenic Assay module overlays individual colonies (A) and calculates the number of colonies (B) in a well.
CHO-K1 cells were treated with a range of Dox concentrations. The formation of colonies over time was analyzed on the Omni platform using the Colony Assay module.
Result: The individual colonies were marked (A) and counted (B) in each well automatically. At these low densities whole-well imaging is crucial for obtaining accurate counts.
Purpose: To establish the timing of the clonogenic assay tipping point. The tipping point identifies the moment in time before colonies get so large they begin to merge, ensuring maximum colony number and size. It is the time point used to calculate the plating efficiency, surviving fraction, and IC50.
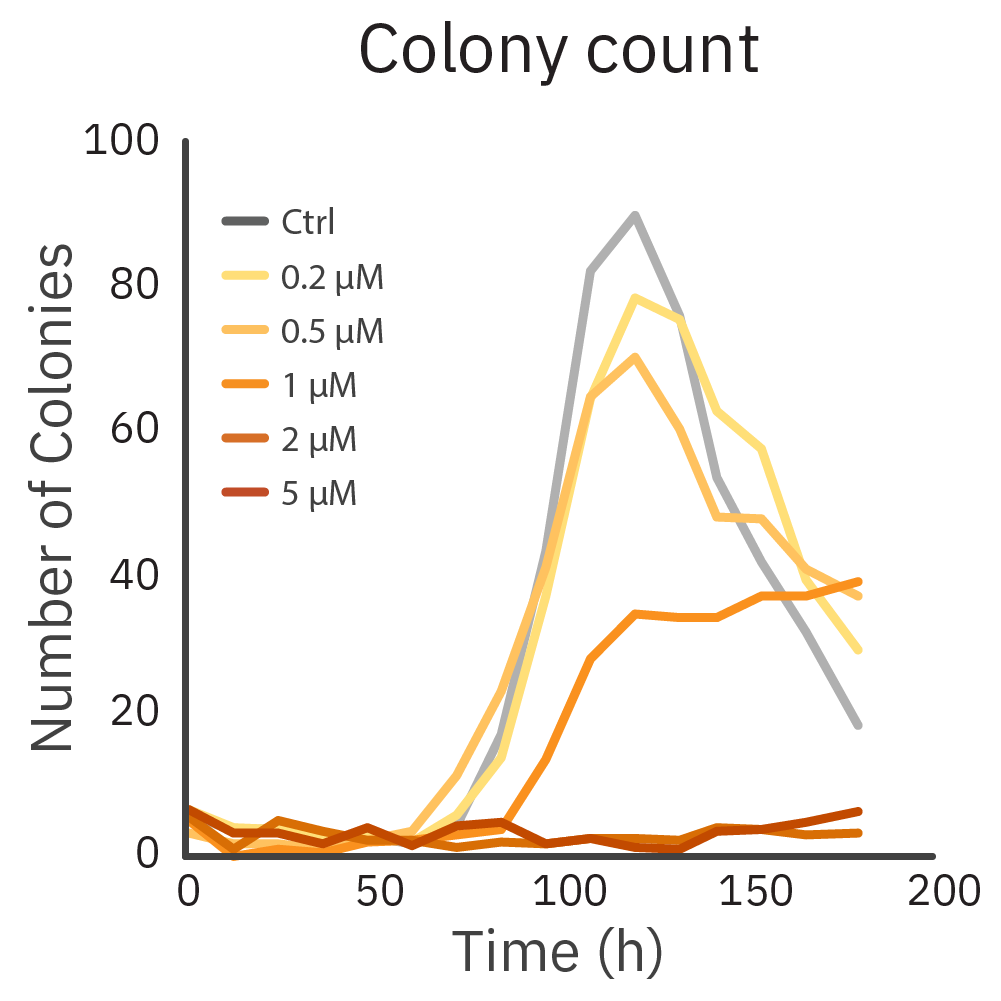
CHO-K1 cells were treated with a range of Dox concentrations. The cells were monitored for 200 hours on the Omni platform to determine a kinetic change in the number of colonies.
Result: The decrease in the number of colonies at 120 hours indicates the assay tipping point.
Purpose: To measure chemotherapeutic response of cancer cells. Clonogenic assays are used to measure reproductive death after therapeutic treatment in many types of cancers.
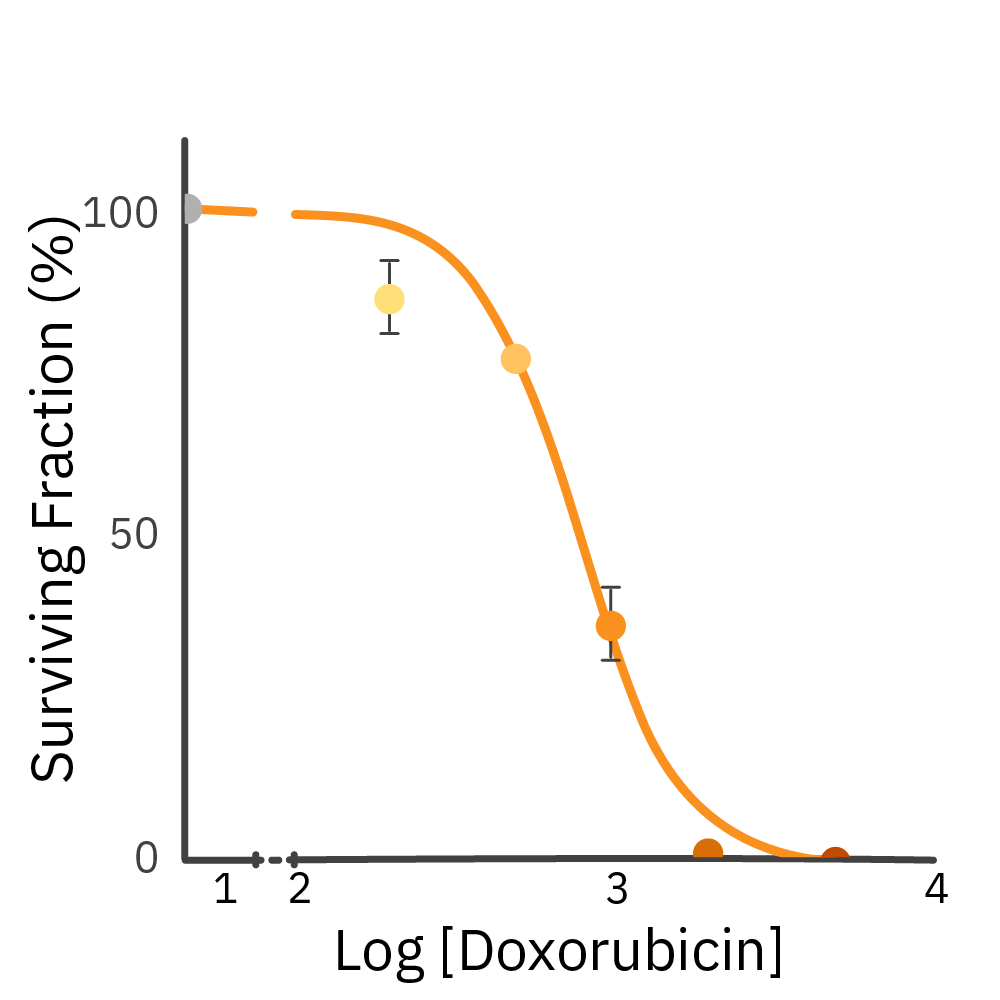
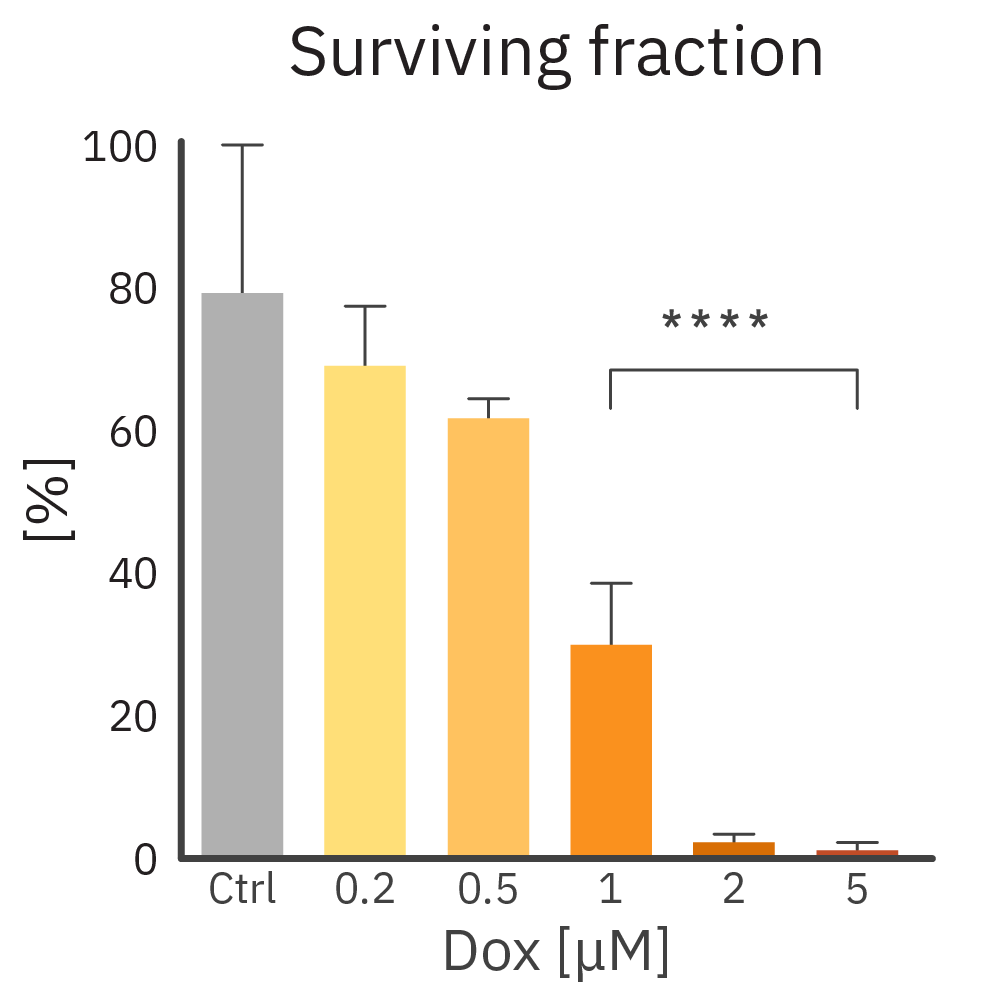
CHO-K1 cells were treated with a chemotherapeutic agent (Dox). Growth was monitored on the Omni platform. Surviving fraction was calculated using the total number of colonies at the culture's tipping point.
Result: Dox inhibits CHO-K1 colony formation in a dose-dependent manner.
Incorporating artificial intelligence (AI) workflows enables users to track colony formation over time and track size, number and circularity automatically.

- >> Accurate – From label-free cell monitoring to fluorescence-based assays, the Omni adds dynamic visual results to any experiment.
- >> Fast – Minimize user-to-user variability with advanced image analysis that incorporates innovative machine-learning algorithms.
- >> Powerful – Get better data with software that assesses the count, size and circularity of colonies in a sample.
Try Omni – hands down the best way to see your cells.
Say goodbye to tedious, time-consuming manual workflows. Complete the form and we’ll show you how automated, AI-powered Omni brings clarity, convenience, and confidence to your cell culturing.



