
Zebrafish are a great translational model of whole organ systems. Their accessibility, ease of genetic engineering, and simple behavioral screens make them useful models for many diseases. Zebrafish models are often used for toxicology and drug discovery research.
Expand on standard zebrafish assays with noninvasive electrophysiological readouts from the brain and spinal cord. Measure real-time drug interactions in live zebrafish. Axion BioSystems’ multielectrode array (MEA) assays are well-suited for studying zebrafish neurobiology.
Scale up organism electrophysiology
-
Measuring neural activity in living zebrafish>
-
Unlock the potential of your zebrafish models with the Maestro MEA>
-
Assay Steps>
Measuring neural activity in living zebrafish has always been a challenge. Consistently capture zebrafish electrophysiology simultaneously across multiple wells from live zebrafish with MEA. Multiple electrodes measure independently across the central nervous system to evaluate neural networks non-invasively and label-free. Specific regions, like the forebrain, midbrain, or hindbrain, may be targeted based on the zebrafish orientation in the well.
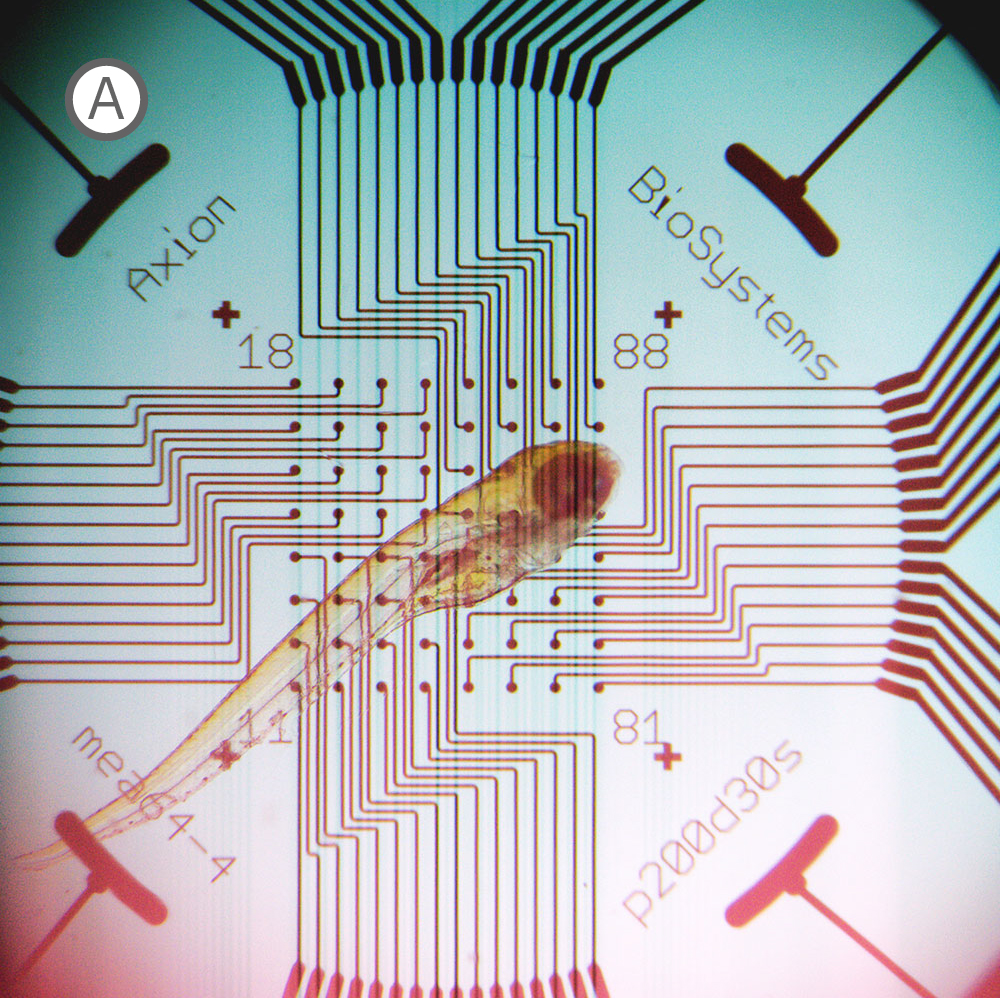

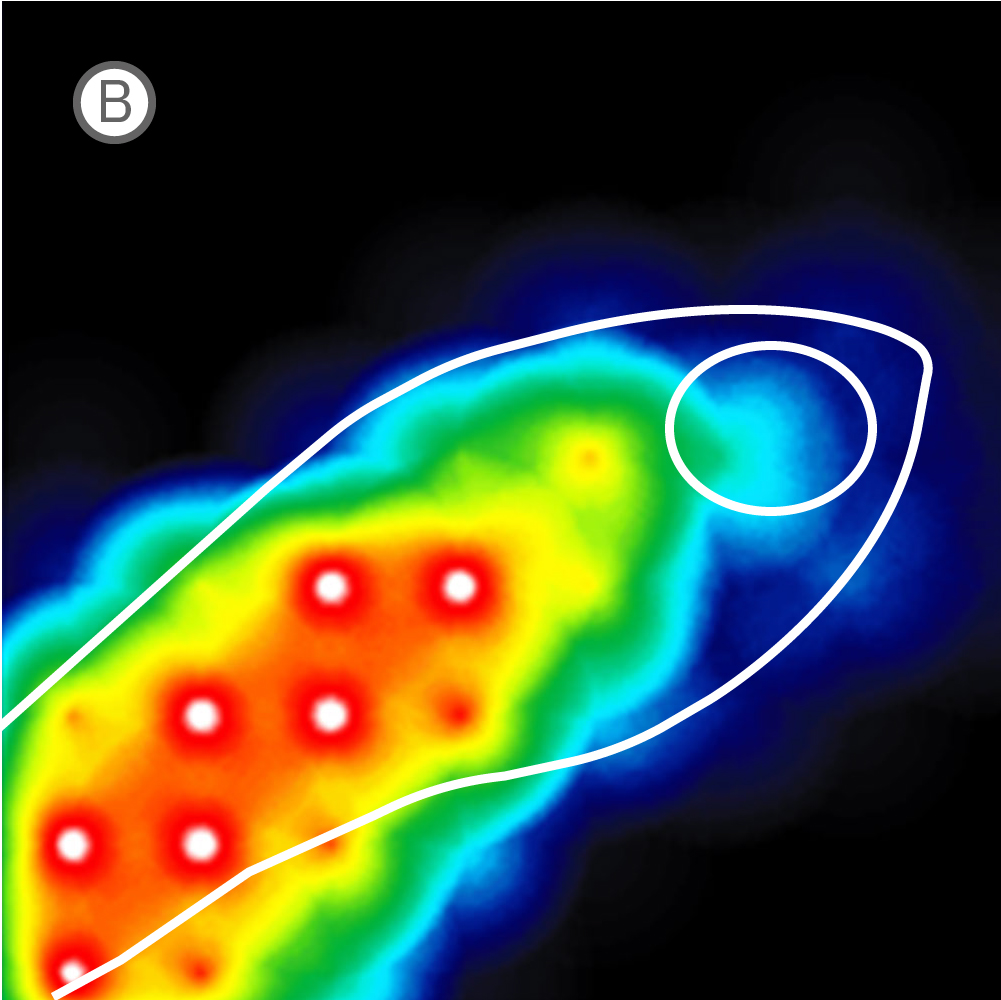
(A) Inverted microscope image of zebrafish larva in a CytoView MEA plate. (B) Neural activity map illustrating that the detected extracellular activity aligns with zebrafish orientation.
The transparent CytoView plates let you identify which electrodes correspond to which region in the spinal cord or brain. With the option to multiplex standard zebrafish imaging assays on CytoView plates after measuring activity, MEA data can easily supplement many workflows. While recording, regions of activity light up bright on the activity map in AxIS Navigator while uncovered electrodes remain dark.
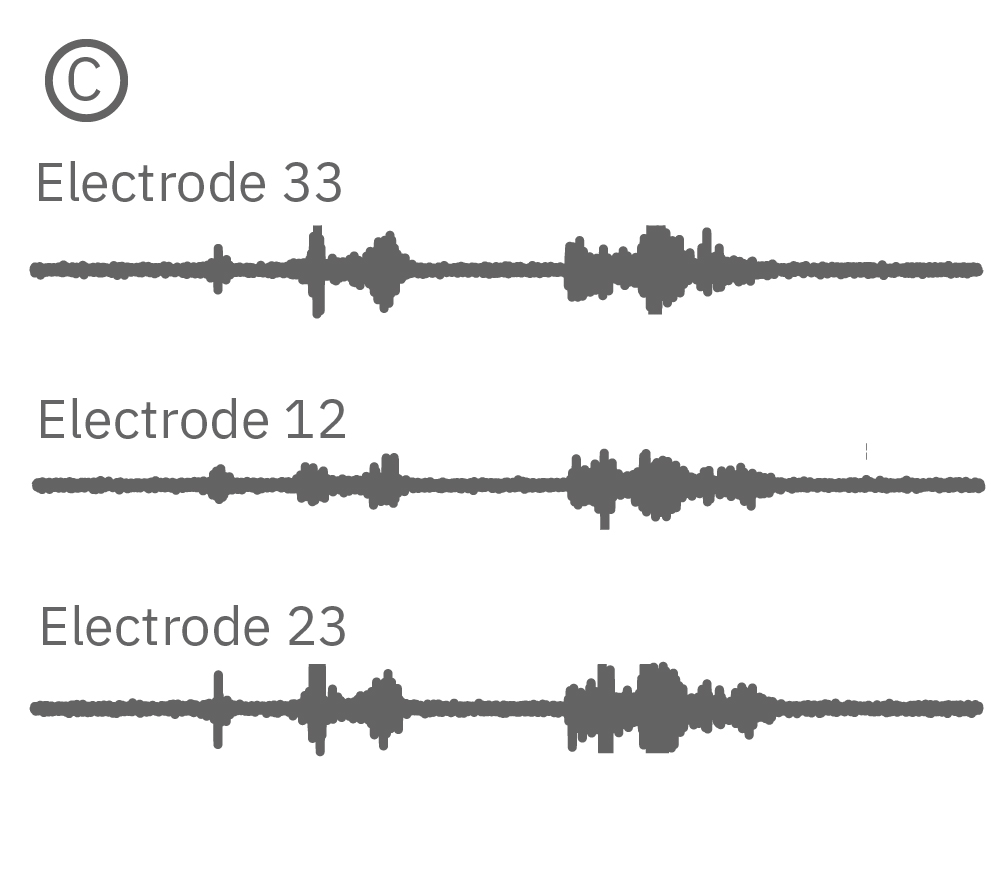
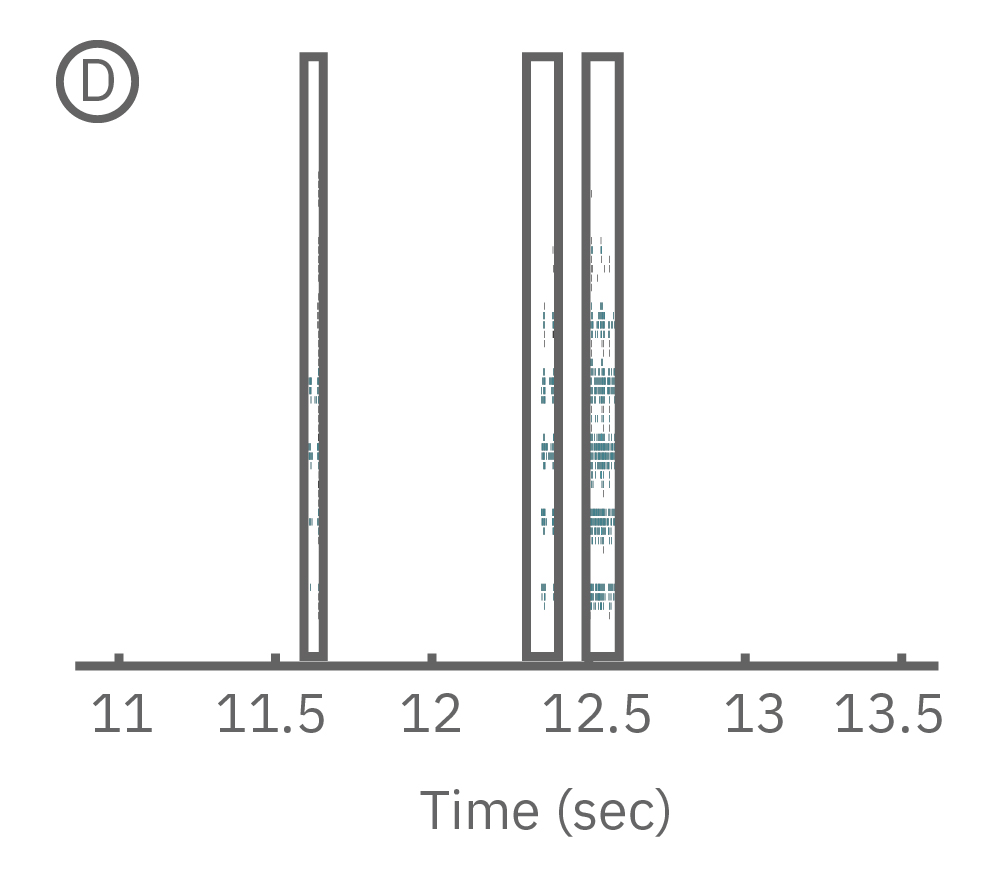
(C) Continuous data recorded from 3 electrodes along a zebrafish spinal cord. (D) Well-wide raster plot demonstrates zebrafish spinal cord activity is organized into synchronous bursts.
AxIS Navigator lets you analyze spiking events from a single electrode, identify and sort individual neurons, or compare between multiple electrodes; allowing you to analyze localized as well as network or region-wide nervous system activity. Zebrafish electrophysiology has never been easier or more accessible than with the Maestro MEA.
With the Maestro MEA, gain new insights into:
- >> Neurodevelopmental and degenerative disorders
- >> Drug or toxicological effects
- >> Simultaneous brain and spinal cord responses
Read the paper below by Tomasello, 2020 to learn more about the step-by-step process to record electrophysiological data from zebrafish on the Maestro MEA system.
Read Tomasello's Zebrafish Publication
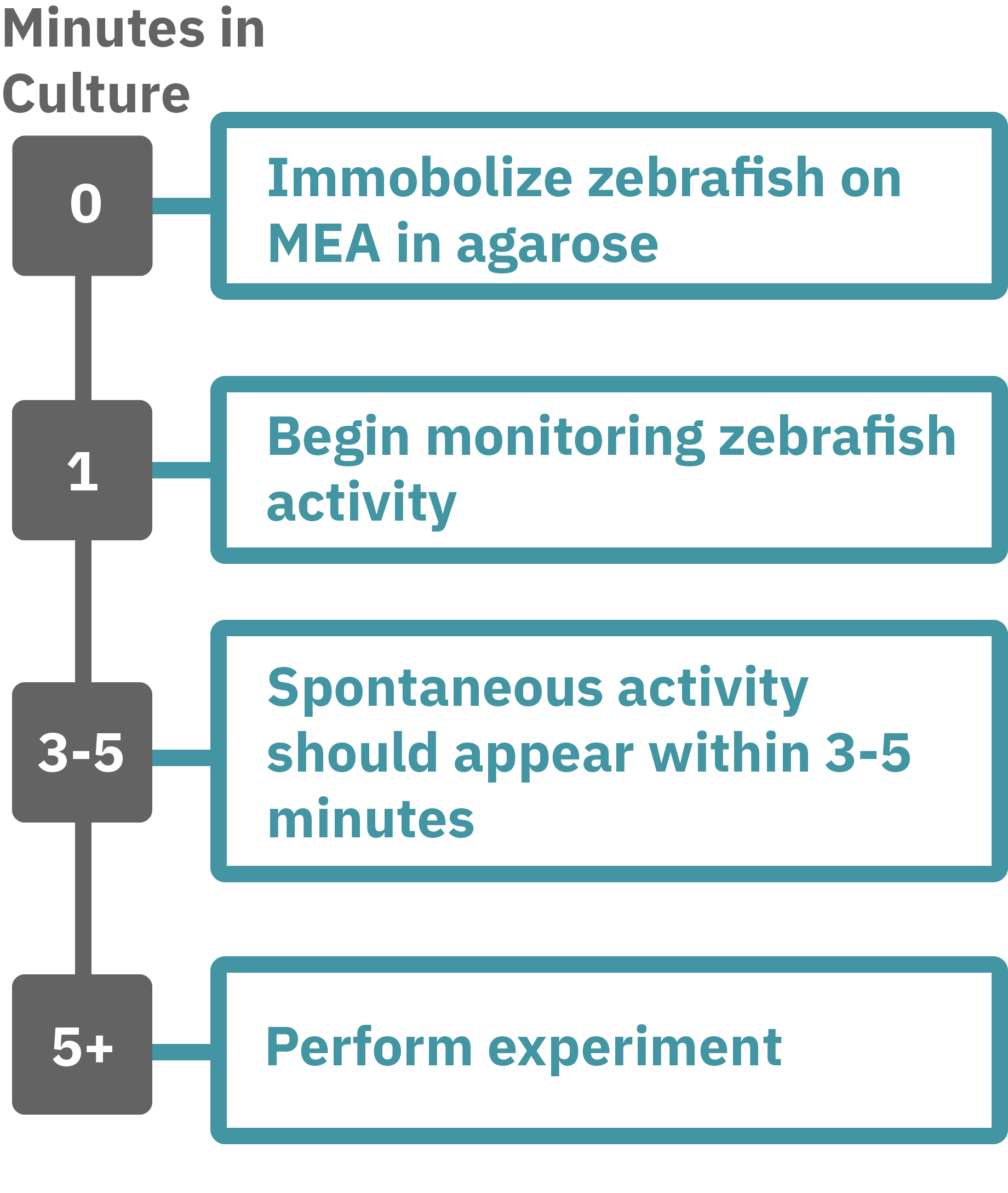
Getting started with Maestro Pro and Edge couldn't be easier. Immobilize the zebrafish in agarose over the electrode containing region of a well in an MEA plate. Once the agarose has solidified, start recording immediately. Spontaneous activity should be detected within 3-5 minutes. Analyze the zebrafish’s neural activity in the MEA plate label-free and in real-time with AxIS Navigator Neural Module software. Add test compounds as required.






